The Model Exploration Software OpenMOLE (Open MOdeL Experiment) explores, diagnoses your numerical model and optimizes its dynamics, taking advantage of distributed computing environments.
The typical usages of OpenMOLE are high performance model calibration, model exploration, machine learning, optimization, data processing, …
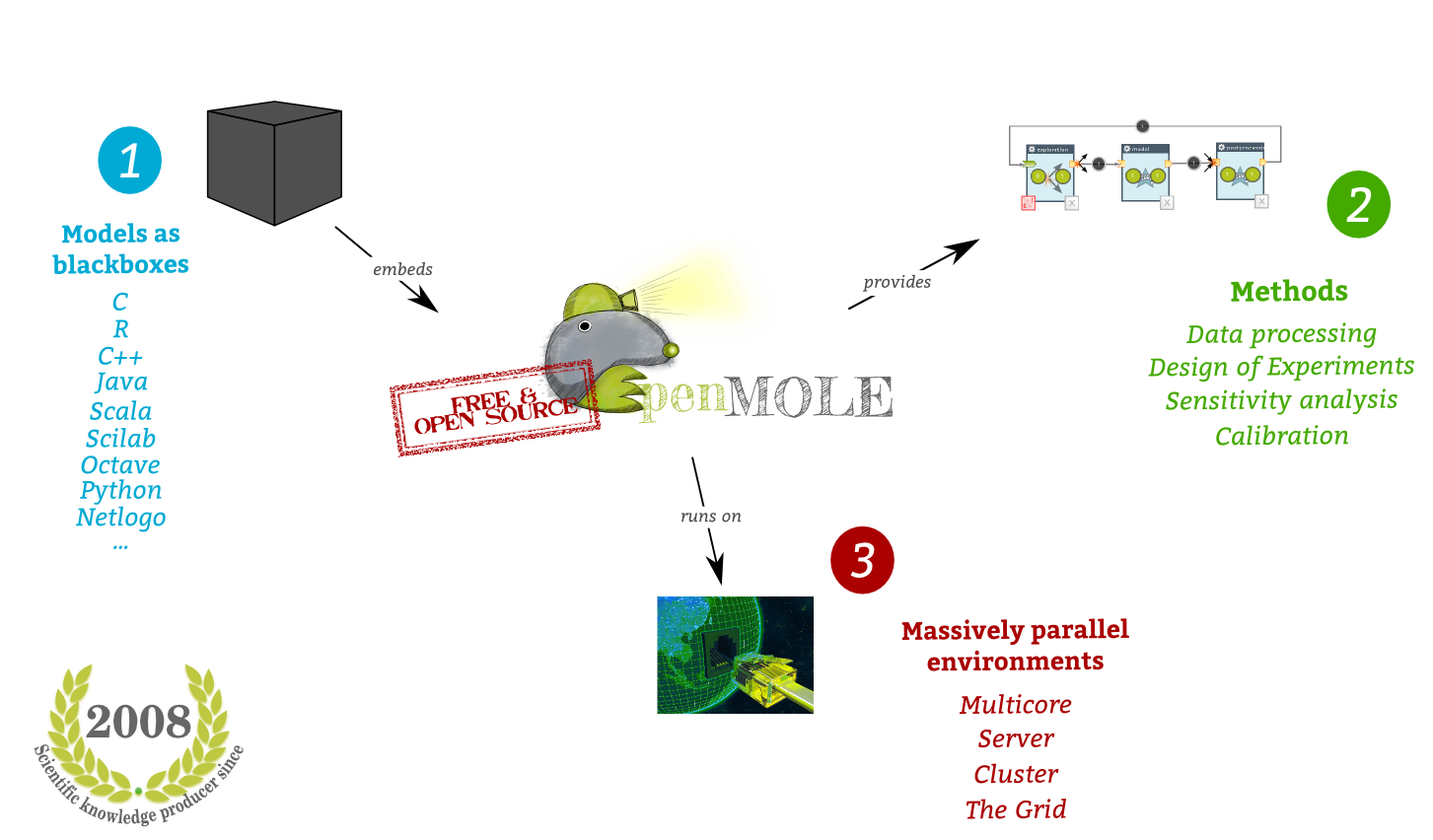
OpenMOLE is:
- Programming language agnostic: it supports any JVM-enabaled language (Java, Scala, NetLogo, …) natively, compiled languages through Linux binaries (C, C++, …), and scripts whose interpreter is available on Linux (R, Python, SciLab, …)
- Designed for Distributed computing - Works on multi-core machines, clusters, grids, desktop grid.
- Expressive – with a workflow language to describe a naturally parallel processes.
- Scalable - Handles millions of tasks, years of computation, and GBs of data.
- Mature - Developed since 2008 and widely used.
- Free and Open Source – under the AGPLv3 free software license.
Related publications:
- Passerat-Palmbach et al., Frontiers Neuroinformatics 2017
- Reuillon et al., HPCS 2015
- Passerat-Palmbach et al., HPC-MICCAI 2014
- Reuillon et al., FGCS 2013
- Reuillon et al., HPCS 2010
This work is done in collaboration with Romain Reuillon and Mathieu Leclaire.